Picritic Basalt¶
XML Input File¶
The basename for this file is basalt_circle.xml.
The file can be run using this command:
microstructpy --demo=basalt_circle.xml
The full text of the file is:
<?xml version="1.0" encoding="UTF-8"?>
<input>
<material>
<name> Plagioclase </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 45.2 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #BDBDBD </color>
</material>
<material>
<name> Olivine </name>
<shape> ellipse </shape>
<fraction>
<dist_type> norm </dist_type>
<loc> 19.1 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> olivine_cdf.csv </filename>
</size>
<aspect_ratio>
<dist_type> uniform </dist_type>
<loc> 1.0 </loc>
<scale> 2.0 </scale>
</aspect_ratio>
<angle_deg>
<dist_type> uniform </dist_type>
<loc> -90 </loc>
<scale> 180 </scale>
</angle_deg>
<color> #99BA73 </color>
</material>
<material>
<name> Diopside </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 13.2 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #709642 </color>
</material>
<material>
<name> Hypersthene </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 16.6 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #876E59 </color>
</material>
<material>
<name> Magnetite </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 3.35 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #6E6E6E </color>
</material>
<material>
<name> Chromite </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 0.65 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>\
</size>
<color> #545454 </color>
</material>
<material>
<name> Ilmenite </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 0.65 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #6B6B6B </color>
</material>
<material>
<name> Apatite </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 3.35 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #ABA687 </color>
</material>
<domain>
<shape> circle </shape>
<diameter> 10 </diameter>
</domain>
<settings>
<directory> basalt_circle </directory>
<filetypes>
<seeds_plot> png </seeds_plot>
<poly_plot> png </poly_plot>
<tri_plot> png </tri_plot>
<verify_plot> png </verify_plot>
</filetypes>
<plot_axes> False </plot_axes>
<verbose> True </verbose>
<verify> True </verify>
<mesh_max_edge_length> 0.01 </mesh_max_edge_length>
<mesh_min_angle> 20 </mesh_min_angle>
<mesh_max_volume> 0.05 </mesh_max_volume>
<seeds_kwargs>
<linewidths> 0.2 </linewidths>
</seeds_kwargs>
<poly_kwargs>
<linewidths> 0.2 </linewidths>
</poly_kwargs>
<tri_kwargs>
<linewidths> 0.1 </linewidths>
</tri_kwargs>
</settings>
</input>
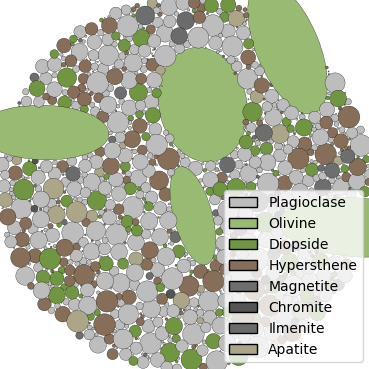
Material 1 - Plagioclase¶
<material>
<name> Plagioclase </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 45.2 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #BDBDBD </color>
</material>
Plagioclase composes approximately 45% of this picritic basalt sample. It is an aphanitic component, meaning fine-grained, and follows a custom size distribution.
Material 2 - Olivine¶
<material>
<name> Olivine </name>
<shape> ellipse </shape>
<fraction>
<dist_type> norm </dist_type>
<loc> 19.1 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> olivine_cdf.csv </filename>
</size>
<aspect_ratio>
<dist_type> uniform </dist_type>
<loc> 1.0 </loc>
<scale> 2.0 </scale>
</aspect_ratio>
<angle_deg>
<dist_type> uniform </dist_type>
<loc> -90 </loc>
<scale> 180 </scale>
</angle_deg>
<color> #99BA73 </color>
</material>
Olivine composes approximately 19% of this picritic basalt sample. There are large phenocrysts of olivine in picritic basalt, so the crystals are generally larger than the other components and have a non-circular shape. The orientation of the phenocrysts is uniform random, with the aspect ratio varying from 1 to 3 uniformly.
Materials 3-8¶
<material>
<name> Diopside </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 13.2 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #709642 </color>
</material>
<material>
<name> Hypersthene </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 16.6 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #876E59 </color>
</material>
<material>
<name> Magnetite </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 3.35 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #6E6E6E </color>
</material>
<material>
<name> Chromite </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 0.65 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>\
</size>
<color> #545454 </color>
</material>
<material>
<name> Ilmenite </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 0.65 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #6B6B6B </color>
</material>
<material>
<name> Apatite </name>
<fraction>
<dist_type> norm </dist_type>
<loc> 3.35 </loc>
<scale> 0.2 </scale>
</fraction>
<size>
<dist_type> cdf </dist_type>
<filename> aphanitic_cdf.csv </filename>
</size>
<color> #ABA687 </color>
</material>
Diopside, hypersthene, magnetite, chromite, ilmenite, and apatie compose approximately 36% of this picritic basalt sample. They are aphanitic components, meaning fine-grained, and follow a custom size distribution.
Domain Geometry¶
<domain>
<shape> circle </shape>
<diameter> 10 </diameter>
</domain>
These materials fill a circular domain with a diameter of 30 mm.
Settings¶
<settings>
<directory> basalt_circle </directory>
<filetypes>
<seeds_plot> png </seeds_plot>
<poly_plot> png </poly_plot>
<tri_plot> png </tri_plot>
<verify_plot> png </verify_plot>
</filetypes>
<plot_axes> False </plot_axes>
<verbose> True </verbose>
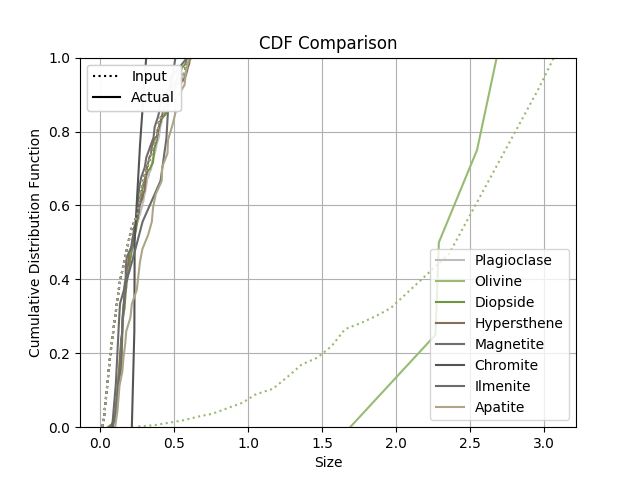
<verify> True </verify>
<mesh_max_edge_length> 0.01 </mesh_max_edge_length>
<mesh_min_angle> 20 </mesh_min_angle>
<mesh_max_volume> 0.05 </mesh_max_volume>
<seeds_kwargs>
<linewidths> 0.2 </linewidths>
</seeds_kwargs>
<poly_kwargs>
<linewidths> 0.2 </linewidths>
</poly_kwargs>
<tri_kwargs>
<linewidths> 0.1 </linewidths>
</tri_kwargs>
</settings>
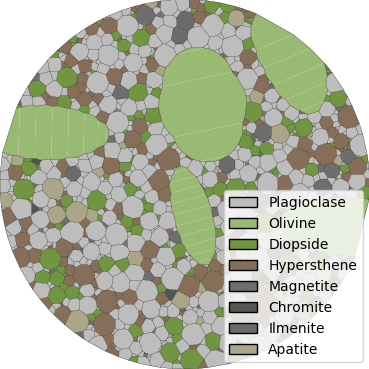
The function will output plots of the microstructure process and those plots
are saved as PNGs.
They are saved in a folder named basalt_circle, in the current directory
(i.e ./basalt_circle).
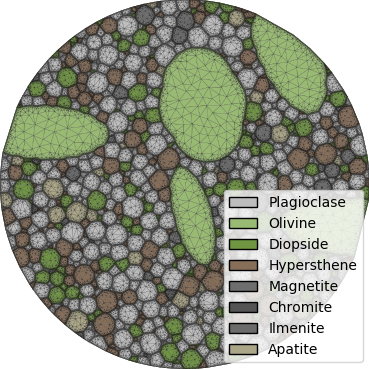
The axes are turned off in these plots, creating PNG files with minimal whitespace.
Mesh controls are introducted to increase grid resolution, particularly at the grain boundaries.